Search
An Easy Assay to Monitor Transfected Levels of the Pre-miR™ miRNA Precursor Positive Control
The Ambion® Pre-miR™ miRNA Starter Kit contains a complete set of reagents needed to begin analyzing miRNA function, including positive and negative control Pre-miR miRNA mimics and siPORT™ NeoFX™ Transfection Agent. When used with the mirVana™ miRNA Isolation Kit or TaqMan® MicroRNA Cells-to-CT™ Kit, a TaqMan MicroRNA Assay can be used to correlate function with delivery of the positive control (Pre-miR hsa-miR-1 miRNA Precursor).
Published research implicates miRNAs in a broad range of diverse biological phenomena, such as cancer, cell growth and development, and viral-host interactions [1–2]. Currently, over 5000 miRNA gene loci are listed in the Sanger Institute database [3]. Functional studies are a critical component in the analysis of a miRNA’s biological activity. In gain-of-function experiments, successful delivery of miRNA mimics (e.g., Pre-miR miRNA Precursors, see sidebar for more information) is crucial for success.
Published research implicates miRNAs in a broad range of diverse biological phenomena, such as cancer, cell growth and development, and viral-host interactions [1–2]. Currently, over 5000 miRNA gene loci are listed in the Sanger Institute database [3]. Functional studies are a critical component in the analysis of a miRNA’s biological activity. In gain-of-function experiments, successful delivery of miRNA mimics (e.g., Pre-miR miRNA Precursors, see sidebar for more information) is crucial for success.
Pre-miR miRNA Starter Kit
The
Pre-miR miRNA Starter Kit enables researchers to deliver functional miRNA mimics into cultured mammalian cells. This kit comprises:
This starter kit enables researchers to optimize transfection for Pre-miR miRNA Precursors into the cell line of choice. The kit components also serve as positive and negative controls in subsequent experiments.
- Pre-miR hsa-miR-1 miRNA Precursor, a positive control
- Pre-miR Negative Control #1
- siPORT NeoFX Transfection Agent
- TaqMan® Gene Expression Assay (Assay ID: Hs00702289_s1) to measure knockdown of Protein Tyrosine Kinase 9 (PTK9) mRNA, the target of the positive control [4]
This starter kit enables researchers to optimize transfection for Pre-miR miRNA Precursors into the cell line of choice. The kit components also serve as positive and negative controls in subsequent experiments.
A Sensitive Measure of Pre-miR Positive Control Levels
The Pre-miR miRNA Starter Kit, combined with methods for RNA preparation that retain small RNAs, such as the
mirVana miRNA Isolation Kit or
TaqMan MicroRNA Cells-to-CT Kit (Figure 1), allows measurement of delivery of the Pre-miR Positive Control with a TaqMan MicroRNA Assay. When the Pre-miR Positive Control is transfected into HeLa cells, a dose-dependent decrease in PTK9 mRNA can be detected using the included TaqMan Gene Expression Assay. Note that introduced levels of hsa-miR-1 can be measured with a corresponding TaqMan MicroRNA Assay (not included).
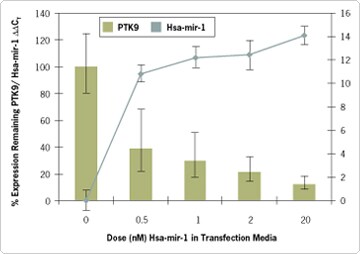
Figure 1. TaqMan® MicroRNA Assays Can be Used to Monitor Levels of Transfected Pre-miR™ miRNA Precursors, While TaqMan Gene Expression Assays Can Monitor Target Downregulation. Methods: HeLa cells (6,000 cells/well) were transfected in 96-well plates with varying doses of either Pre-miR hsa-mir-1 miRNA Precursor or Pre-miR Negative Control #1 using siPORT™ NeoFX™ Transfection Agent (0.3 µL). Media were changed 24 hrs post-transfection, and cells were harvested 48 hrs post-transfection. Total RNA including small RNA was isolated with mirVana™ miRNA Isolation Kit, or cell lysates were prepared with TaqMan Gene Expression Cells-to-CT™ Kit. Quantitative RT-PCR Analysis: TaqMan Gene Expression Assays and TaqMan MicroRNA Assays showed a ΔΔCT shift in PTK9 mRNA expression corresponding to a reduction in PTK9 mRNA expression concurrent with the measurable increase in hsa-mir-1 miRNA mimic levels. PTK9 mRNA CT values were normalized using CT values for 18S rRNA. Hsa-miR-1 miRNA CT values were normalized using RNU44 snoRNA. Hsa-mir-1 is not endogenously expressed in HeLa cells at detectable levels; thus, upregulation of Hsa-mir-1 is presented as |ΔΔCT|. Relative gene expression was calculated using the Comparative CT Method [5].
The Pre-miR miRNA Starter Kit, used in conjunction with TaqMan MicroRNA Assays, provides the researcher with powerful tools to optimize transfection and methods to assess miRNA mimic delivery.
Scientific Contributors
Tera Schaller, Sarah LaMartina, Joe Krebs*, Laura Chapman, and Susan Magdaleno • Applied Biosystems, Austin, TX *Currently at BIOO Scientific Corporation, Austin, TX

Figure 1. TaqMan® MicroRNA Assays Can be Used to Monitor Levels of Transfected Pre-miR™ miRNA Precursors, While TaqMan Gene Expression Assays Can Monitor Target Downregulation. Methods: HeLa cells (6,000 cells/well) were transfected in 96-well plates with varying doses of either Pre-miR hsa-mir-1 miRNA Precursor or Pre-miR Negative Control #1 using siPORT™ NeoFX™ Transfection Agent (0.3 µL). Media were changed 24 hrs post-transfection, and cells were harvested 48 hrs post-transfection. Total RNA including small RNA was isolated with mirVana™ miRNA Isolation Kit, or cell lysates were prepared with TaqMan Gene Expression Cells-to-CT™ Kit. Quantitative RT-PCR Analysis: TaqMan Gene Expression Assays and TaqMan MicroRNA Assays showed a ΔΔCT shift in PTK9 mRNA expression corresponding to a reduction in PTK9 mRNA expression concurrent with the measurable increase in hsa-mir-1 miRNA mimic levels. PTK9 mRNA CT values were normalized using CT values for 18S rRNA. Hsa-miR-1 miRNA CT values were normalized using RNU44 snoRNA. Hsa-mir-1 is not endogenously expressed in HeLa cells at detectable levels; thus, upregulation of Hsa-mir-1 is presented as |ΔΔCT|. Relative gene expression was calculated using the Comparative CT Method [5].
The Pre-miR miRNA Starter Kit, used in conjunction with TaqMan MicroRNA Assays, provides the researcher with powerful tools to optimize transfection and methods to assess miRNA mimic delivery.
Scientific Contributors
Tera Schaller, Sarah LaMartina, Joe Krebs*, Laura Chapman, and Susan Magdaleno • Applied Biosystems, Austin, TX *Currently at BIOO Scientific Corporation, Austin, TX
Ambion® Pre-miR™ miRNA PrecursorsPre-miR™ miRNA Precursors are small, chemically modified, double-stranded RNA molecules designed to mimic endogenous, mature miRNA molecules when transfected or electroporated into cells. These miRNA mimics enable functional analysis by up-regulation of miRNA activity. Examples of experimental designs include miRNA target site identification and validation, screening for miRNAs that regulate expression of a gene, and screening for miRNAs that affect a cellular process.
A potent positive control (targeting hsa-miR-1), non-specific negative controls (some with Cy™3 or FAM™ dye labels), and libraries of Pre-miR Precursors are also available.
MicroRNA-related cellular functions identified through use of Pre-miR miRNA Precursors can be further investigated using Anti-miR™ miRNA Inhibitors, which enable loss-of-function experiments through reduction of miRNA activity.
A potent positive control (targeting hsa-miR-1), non-specific negative controls (some with Cy™3 or FAM™ dye labels), and libraries of Pre-miR Precursors are also available.
MicroRNA-related cellular functions identified through use of Pre-miR miRNA Precursors can be further investigated using Anti-miR™ miRNA Inhibitors, which enable loss-of-function experiments through reduction of miRNA activity.
Pre-miR miRNA Precursors in ActionMicroRNAs Alter Response to Cancer Therapy
A recent study by Weidhass and colleagues examined how miRNA levels affect the response of cancer cells to radiation [1]. They discovered that the let-7 family of miRNAs is overrepresented in the miRNA population with radiation-responsive expression levels. In addition, a radiosensitive state was created when select let-7 family members were overexpressed in vitro in lung cancer cells (using Pre-miR miRNA Precursors) and in vivo in a C. elegans model of radiation-induced cell death, whereas decreasing their levels causes radioresistance. The significance of the let-7 family in the radiation response is further highlighted by showing that altering let-7 levels before radiation affected cellular survival.
1. Weidhaas JB, Babar I, Nallur SM, Trang P, Roush S, Boehm M, Gillespie E, and Slack FJ (2007) MicroRNAs as potential agents to alter resistance to cytotoxic anticancer therapy. Cancer Res 67(23):11111–11116.
A recent study by Weidhass and colleagues examined how miRNA levels affect the response of cancer cells to radiation [1]. They discovered that the let-7 family of miRNAs is overrepresented in the miRNA population with radiation-responsive expression levels. In addition, a radiosensitive state was created when select let-7 family members were overexpressed in vitro in lung cancer cells (using Pre-miR miRNA Precursors) and in vivo in a C. elegans model of radiation-induced cell death, whereas decreasing their levels causes radioresistance. The significance of the let-7 family in the radiation response is further highlighted by showing that altering let-7 levels before radiation affected cellular survival.
1. Weidhaas JB, Babar I, Nallur SM, Trang P, Roush S, Boehm M, Gillespie E, and Slack FJ (2007) MicroRNAs as potential agents to alter resistance to cytotoxic anticancer therapy. Cancer Res 67(23):11111–11116.
References1. Calin GA and Croce CM (2006) MicroRNA signatures in human cancers. Nat Rev Cancer 6:857–866.
2. Sullivan CS and Ganem D (2005) MicroRNAs and viral infection. Mol Cell 20:3–7.
3. Sanger Institute miRNA database; http://microrna.sanger.ac.uk/
sequences.
4. Lim LP, Lau NC, Garrett-Engele P, Grimson A, Schelter JM, Castle J, Bartel DP, Linsley PS, and Johnson JM (2005) Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature 433:769–773.
5. Guide to Performing Relative Quantitation of Gene Expression Using Real-Time Quantitative PCR. Publication #4303859, available at www.appliedbiosystems.com.
2. Sullivan CS and Ganem D (2005) MicroRNAs and viral infection. Mol Cell 20:3–7.
3. Sanger Institute miRNA database; http://microrna.sanger.ac.uk/
sequences.
4. Lim LP, Lau NC, Garrett-Engele P, Grimson A, Schelter JM, Castle J, Bartel DP, Linsley PS, and Johnson JM (2005) Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature 433:769–773.
5. Guide to Performing Relative Quantitation of Gene Expression Using Real-Time Quantitative PCR. Publication #4303859, available at www.appliedbiosystems.com.