Search
Fluorescence Polarization (FP)—Note 1.4
Principles
Fluorescence polarization measurements provide information on molecular orientation and mobility and processes that modulate them, including receptor–ligand interactions, protein–DNA interactions, proteolysis, membrane fluidity and muscle contraction (Figure 1).
Because polarization is a general property of fluorescent molecules (with certain exceptions such as lanthanide chelates), polarization-based readouts are somewhat less dye dependent and less susceptible to environmental interferences such as pH changes than assays based on fluorescence intensity measurements. Experimentally, the degree of polarization is determined from measurements of fluorescence intensities parallel and perpendicular with respect to the plane of linearly polarized excitation light, and is expressed in terms of fluorescence polarization (P) or anisotropy (r):

Note that both P and r are ratio quantities with no nominal dependence on dye concentration. Because of the ratio formulation, fluorescence intensity variations due to the presence of colored sample additives tend to cancel and produce relatively minor interferences.![]() P has physically possible values ranging from –0.33 to 0.5. In practice, these limiting values are rarely attained. Measured values of P in bioanalytical applications typically range from 0.01 to 0.3 or 10 to 300 mP (mP = P/1000). This measurement range is not as narrow as it might appear to be because very precise measurements (P ± 0.002 or ± 2 mP) are readily obtainable with modern instrumentation.
P has physically possible values ranging from –0.33 to 0.5. In practice, these limiting values are rarely attained. Measured values of P in bioanalytical applications typically range from 0.01 to 0.3 or 10 to 300 mP (mP = P/1000). This measurement range is not as narrow as it might appear to be because very precise measurements (P ± 0.002 or ± 2 mP) are readily obtainable with modern instrumentation.
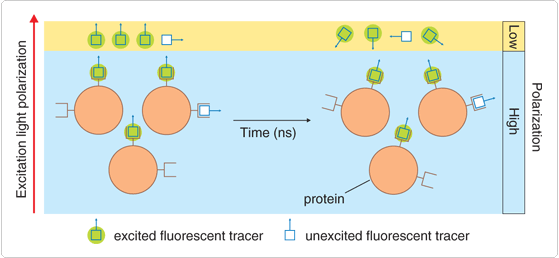
Figure 1. Physical basis of fluorescence polarization assays. Dye molecules with their absorption transition vectors (arrows) aligned parallel to the electric vector of linearly polarized light (along the vertical page axis) are selectively excited. For dyes attached to small, rapidly rotating molecules, the initially photoselected orientational distribution becomes randomized prior to emission, resulting in low fluorescence polarization. Conversely, binding of the low molecular weight tracer to a large, slowly rotating molecule results in high fluorescence polarization. Fluorescence polarization therefore provides a direct readout of the extent of tracer binding to proteins, nucleic acids and other biopolymers.
Dependence of Fluorescence Polarization on Molecular Mobility
Interpretation of the dependence of fluorescence polarization on molecular mobility is usually based on a model derived in 1926 from the physical theory of Brownian motion by Perrin:![]()
![]()
where Po is the fundamental polarization of the dye (for fluorescein, rhodamine and BODIPY dyes, Po is close to the theoretical maximum of 0.5), τ is the excited-state lifetime of the dye and Φ is the rotational correlation time of the dye or dye conjugate. These relationships can be expressed in terms of fluorescence anisotropy in an equivalent and mathematically simpler manner. For a hydrodynamic sphere, Φ can be estimated as follows:
![]()
where η = solvent viscosity, T = temperature, R = gas constant and V = molecular volume of the fluorescent dye or dye conjugate. In turn, V can be estimated from the molecular weight of the dye or dye conjugate with appropriate adjustments for hydration. Simulations of these relationships are shown in Figure 2., leading to the following general conclusions:
- Fluorescence polarization increases as molecular weight increases.
- Fluorescence polarization increases as solvent viscosity increases.
- Fluorescence polarization decreases as the excited state lifetime of the dye (τ) increases.
Note that these simulations assume that the dye is rigidly attached to a spherical carrier. When conventional parameter estimates for proteins in aqueous solutions are used, Φ is found to increase by about 1 ns per 2400 dalton increase of molecular weight.![]()
 | Figure 2. Simulation of the relationship between molecular weight (MW) and fluorescence polarization (P). Simulations are shown for dyes with various fluorescence lifetimes (τ): 1 ns (cyanine dyes) in purple, 4 ns (fluorescein and Alexa Fluor 488 dyes) in red, 6 ns (some BODIPY dyes) in green and 20 ns (dansyl dyes) in blue. At MW = 1000, P = 0.167 for τ = 1 ns, P = 0.056 for τ = 4 ns, P = 0.039 for τ = 6 ns and P = 0.012 for τ = 20 ns. Simulations assume Po (the fundamental polarization) = 0.5 and rigid attachment of dyes to spherical carriers. |
Dyes for Fluorescence Polarization Assays
Tracers used in fluorescence polarization assays include peptides, drugs and cytokines that are modified by the attachment of a fluorescent dye. Depolarization due to flexibility in the attachment of the dye, sometimes referred to as the "propeller effect," distorts the relationships between P and molecular weight shown in Figure 2. For this reason, it is generally preferable to use reactive dyes without aliphatic linkers between the fluorophore and the reactive group in the preparation of tracers for fluorescence polarization assays.![]()
A key factor in the performance of fluorescence polarization assays is the extent to which the biological activity of the tracer is perturbed by the dye modification. BODIPY dyes generally produce less perturbation of receptor-binding affinity and other activity parameters than conventional dyes such as fluorescein and rhodamine.![]() Furthermore, BODIPY dyes usually have longer excited-state lifetimes than fluorescein and rhodamine dyes, making their fluorescence polarization sensitive to binding interactions over a larger molecular weight range (Figure 2). The long-wavelength BODIPY TMR and BODIPY TR dyes also tend to minimize assay interferences due to intrinsically fluorescent sample contaminants.
Furthermore, BODIPY dyes usually have longer excited-state lifetimes than fluorescein and rhodamine dyes, making their fluorescence polarization sensitive to binding interactions over a larger molecular weight range (Figure 2). The long-wavelength BODIPY TMR and BODIPY TR dyes also tend to minimize assay interferences due to intrinsically fluorescent sample contaminants.![]()
Applications
Fluorescence polarization measurements have long been a valuable biophysical research tool for investigating processes such as membrane lipid mobility, myosin reorientation and protein–protein interactions at the molecular level.![]() Immunoassays that have been developed and used extensively for clinical diagnostics represent the largest group of bioanalytical applications.
Immunoassays that have been developed and used extensively for clinical diagnostics represent the largest group of bioanalytical applications.![]() The more recent advent of microplate readers equipped with polarizing optics has led to the adoption of fluorescence polarization as a readout mode for high-throughput screening.
The more recent advent of microplate readers equipped with polarizing optics has led to the adoption of fluorescence polarization as a readout mode for high-throughput screening.![]() Some typical bioanalytical applications of fluorescence polarization–based assays are summarized below (Table 1).
Some typical bioanalytical applications of fluorescence polarization–based assays are summarized below (Table 1).
Table 1. Examples of fluorescence polarization assays
| Assay Target | Tracer | References |
|---|---|---|
| Ligand binding to neurokinin 1 (NK1) receptor | Fluorescein-labeled substance P | Biochemistry (1994) 33:13079 |
| Ligand binding to melanocortin G-protein–coupled receptors | BODIPY TMR dye–labeled NDP-αMSH | J Biomol Screen (2000) 5:329 |
| Ligand binding to B2 bradykinin receptor, a G-protein–coupled receptor | BODIPY TMR dye–labeled HOE140 | J Biomol Screen (2002) 7:111 |
| Ligand binding to estrogen receptors | Fluorescein-labeled estradiol | J Biomol Screen (2000) 5:77 |
| Ligand binding to tyrosine kinase Src homology domains | Fluorescein- and BODIPY TR dye–labeled phosphopeptides | Anal Biochem (1999) 275:62; Anal Biochem (1997) 247:77 |
| Substrate binding to protein farnesyltransferase | Oregon Green 488 dye–labeled peptide | Biochemistry (1999) 38:13138 |
| β-Lactam antibiotic binding to penicillin-binding proteins | BODIPY FL dye–labeled penicillin V | Antimicrob Agents Chemother (1999) 43:1124 |
| Protein kinase activity | Fluorescently labeled phosphopeptide | Anal Biochem (2000) 278:206; Methods (2000) 22:61 |
| Nonspecific protease activity | BODIPY FL dye–labeled casein | Anal Biochem (1996) 243:1 |
| Detection of specific PCR products | Fluorescein-labeled oligonucleotide | Gene (2000) 259:123 |
| Ligation and cleavage of RNA by ribozymes | Fluorescein- or tetramethylrhodamine-labeled oligoribonucleotide | Biotechniques (2000) 29:344 |
| SNP detection by allele-specific primer extension | Fluorescent ddNTP | Genome Res (1999) 9:492 |
| Protein–protein and protein–nucleic acid interactions | Alexa Fluor 488 dye–labeled human Factor VIIa, Oregon Green 488 dye–labeled soluble human tissue factor and Oregon Green 514 dye–labeled oligonucleotide | Anal Biochem (2002) 308:18 |
| Oligomerization and fibril formation of α-synuclein | Oregon Green 488 dye– and Alexa Fluor 594 dye–labeled α-synuclein | Biochemistry (2007) 46:12522 |
For Research Use Only. Not for use in diagnostic procedures.