Search
Using Organic Fluorescent Probes in Combination with GFP—Note 12.1
Probes for Multiplexed Detection of GFP-Expressing Cells
The Green Fluorescent Protein (GFP) reporter has added a new dimension to the analysis of protein localization, allowing real-time examination in live cells of processes that have conventionally been observed through immunocytochemical "snapshots" in fixed specimens.![]() Using spectrally distinct, organic fluorescent probes and markers (Table 1) adds extra data dimensions and reference points to these experiments (Figure 1).
Using spectrally distinct, organic fluorescent probes and markers (Table 1) adds extra data dimensions and reference points to these experiments (Figure 1).
 | Figure 1. The morphology of sporulating Bacillus subtilis in the early stages of forespore engulfment. The membranes and chromosomes of both the forespore and the larger mother cell are stained with FM 4-64 (red; T3166, T13320) and DAPI (blue; D1306, D3571, D21490), respectively. The small green-fluorescent patch indicates the localization of a GFP fusion to SPoIIIE, a protein essential for translocation of the forespore chromosome that may also regulate membrane fusion events (see Proc Natl Acad Sci U S A (1999) 96:14553). The background contains sporangia at various stages in the engulfment process stained with MitoTracker Green FM (green, M7514) and FM 4-64 (red). |
The majority of the applications summarized in Table 1 involve live cells, tissues and organisms. There are many other instances where research objectives call for complementary use of immunochemical and GFP-based protein localization techniques. These experiments demand the combination of brightness, photostability and spectral separation provided by our Alexa Fluor dye–labeled secondary detection reagents. For two-color combinations with GFP, we recommend the Alexa Fluor 555, Alexa Fluor 568 or Alexa Fluor 594 dye–labeled secondary antibodies (Secondary Immunoreagents—Section 7.2, Summary of Molecular Probes secondary antibody conjugates—Table 7.1). The addition of Alexa Fluor 635 or Alexa Fluor 647 dye–labeled antibodies allows three-color detection. Some immunohistochemical procedures such as paraffin embedding of fixed tissue result in loss of the intrinsic fluorescence of GFP. In other cases, GFP expression levels may simply be too low for detection above background autofluorescence.![]() Antibodies to GFP provide remedies for these problems (Figure 2). We offer unlabeled mouse and rabbit monoclonal GFP antibodies and unlabeled rabbit and chicken polyclonal GFP antibodies, as well as Alexa Fluor dye–labeled rabbit polyclonal GFP antibodies (Antibodies against Expression Tags—Section 7.5).
Antibodies to GFP provide remedies for these problems (Figure 2). We offer unlabeled mouse and rabbit monoclonal GFP antibodies and unlabeled rabbit and chicken polyclonal GFP antibodies, as well as Alexa Fluor dye–labeled rabbit polyclonal GFP antibodies (Antibodies against Expression Tags—Section 7.5).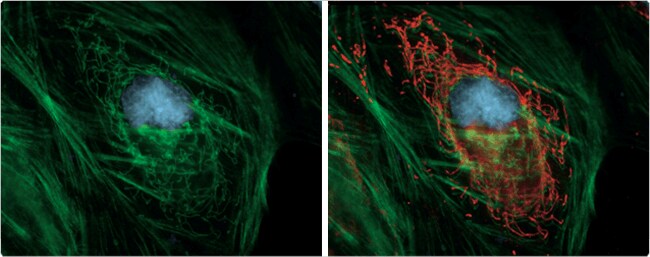
Figure 2. HeLa cell transfected with pShooter pCMV/myc/mito/GFP, then fixed and permeabilized. Green Fluorescent Protein (GFP) localized in the mitochondria was labeled with mouse IgG2a anti-GFP antibody (A11120) and detected with orange-fluorescent Alexa Fluor 555 goat anti–mouse IgG antibody (A21422), which colocalized with the dim GFP fluorescence. F-actin was labeled with green-fluorescent Alexa Fluor 488 phalloidin (A12379), and the nucleus was stained with blue-fluorescent DAPI (D1306, D3571, D21490). The sample was mounted using ProLong Gold antifade reagent (P36930). Some GFP fluorescence is retained in the mitochondria after fixation (left), but immunolabeling and detection greatly improve visualization (right).
Alexa Fluor Dyes: Highly Fluorescent FRET Acceptors
Proximity-dependent fluorescence resonance energy transfer (FRET) allows detection of protein–protein interactions with much higher spatial resolution than conventional diffraction-limited microscopy.![]() Alexa Fluor dyes with strong absorption in the 500–600 nm wavelength range are excellent FRET acceptors from GFP (Table 2). An assay to detect activation of GFP–GTPase fusions developed by researchers at Scripps Research Institute
Alexa Fluor dyes with strong absorption in the 500–600 nm wavelength range are excellent FRET acceptors from GFP (Table 2). An assay to detect activation of GFP–GTPase fusions developed by researchers at Scripps Research Institute ![]() utilizes the GTPase-binding domain (PBD) of PAK1, a protein that binds to GTPases only in their activated GTP-bound form. GTPase activation is indicated by FRET from GFP to PDB labeled with Alexa Fluor 546 C5-maleimide at a single N-terminal cysteine residue. This assay has been used to determine the location and dynamics of rac and Cdc42 GTPase activation in live cells.
utilizes the GTPase-binding domain (PBD) of PAK1, a protein that binds to GTPases only in their activated GTP-bound form. GTPase activation is indicated by FRET from GFP to PDB labeled with Alexa Fluor 546 C5-maleimide at a single N-terminal cysteine residue. This assay has been used to determine the location and dynamics of rac and Cdc42 GTPase activation in live cells.![]()
Normalizing Expression and Translation Signals
In 2002, researchers in Scott Fraser's laboratory at the California Institute of Technology reported a method of coinjecting Texas Red dye–labeled 10,000 MW dextran and GFP vectors into sea urchin embryos. This method overcomes a multitude of problems inherent in making intra- and inter-embryo comparisons of gene expression levels using confocal microscopy. In particular, laser excitation and fluorescence collection efficiencies vary with the depth of the fluorescent protein in the embryo, and the orientation of different embryos on the coverslip varies relative to the microscope objective. Measuring the ratio of GFP fluorescence to Texas Red dextran fluorescence corrects for these spatial factors, providing a gene expression readout that is 2–50 times more accurate than conventional confocal microscopy procedures depending on the localization of GFP within an embryo.![]() A similar strategy was previously used to determine translation efficiencies of GFP-encoding mRNAs.
A similar strategy was previously used to determine translation efficiencies of GFP-encoding mRNAs.![]()
Table 1. Probes for multiplexed detection of GFP-expressing cells.*
| Target | Probe | Cat. No. | Ex/Em † | GFP Fusion Partner | Specimen | Reference | |
|---|---|---|---|---|---|---|---|
| Physiological Indicators | |||||||
| Intracellular Ca2+ | Fura-2 AM | F1201, F1221, F1225, F14185 | 335/505 ‡ | Protein kinase C (PKC) | BHK cells | Biochem J (1999) 337:211 | |
| Intracellular Ca2+ | X-Rhod-1 AM | X14210 | 580/602 | Trpm5 (melastatin-related cation channel) | CHO cells | Nat Neurosci (2002) 5:1169 | |
| Intracellular Ca2+ | Fura Red AM | F3020, F3021 | 488/650 | GFP expressed specifically in pancreatic β-cells | Mouse pancreatic islets | Am J Physiol Endocrinol Metab (2003) 284:E177 | |
| Intracellular pH | 5-(and 6-)Carboxy SNARF-1 AM ester acetate | C1271 | 568/635 | Human growth hormone (hGH) | RIN1046-38 insulinoma cells | Am J Physiol Cell Physiol (2002) 283:C429 | |
| Mitochondrial membrane potential | TMRM | T668 | 555/580 | Cytochrome c | MCF-7 human breast carcinoma, HeLa | J Cell Sci (2003) 116:525 | |
| Superoxide (O2–) | Dihydroethidium | D1168 | 518/605 | Cytochrome c | MCF-7 human breast carcinoma | J Biol Chem (2003) 278:12645 | |
| Synaptic activity | FM 4-64 | T3166, T13320 | 506/750 § | VAMP (vesicle-associated membrane protein) | Rat hippocampal neurons | Nat Neurosci (2000) 3:445 | |
| Receptors and Endocytosis | |||||||
| Acetylcholine receptor | Tetramethylrhodamine α-bungarotoxin | T1175 | 553/577 | Rapsyn (receptor-aggregating protein) | Zebrafish | J Neurosci (2001) 21:5439 | |
| Epidermal growth factor (EGF) | Rhodamine EGF | E3481 | 555/581 | EGF receptor | MTLn3 rat mammary adenocarcinoma | Mol Biol Cell (2000) 11:3873 | |
| Endosomes | Transferrin from human serum, Alexa Fluor 546 conjugate | T23364 | 556/573 | β2-adrenergic receptor (β2AR) | HEK 293, rat hippocampal neurons | Brain Res (2003) 984:21 | |
| Endosomes | Transferrin from human serum, Alexa Fluor 568 conjugate | T23365 | 578/603 | PrPc (cellular prion protein) | SN56 cells | J Biol Chem (2002) 277:33311 | |
| Endosomes | FM 4-64 | T3166, T13320 | 506/750 § | PrPc (cellular prion protein) | SN56 cells | J Biol Chem (2002) 277:33311 | |
| Organelles | |||||||
| Endoplasmic reticulum | ER-Tracker Blue-White DPX | E12353 | 375/520 ‡ | HSD17B7 gene product (3-ketosteroid reductase) | HeLa, NIH 3T3 | Mol Endocrinol (2003) 17:1715 | |
| Golgi complex | BODIPY TR ceramide | D7540 | 589/617 | PrPc (cellular prion protein) | SN56 cells | J Biol Chem (2002) 277:33311 | |
| Lysosomes | LysoTracker Red | L7528 | 577/590 | Heparanase | Primary human fibroblasts, MDA-231 (human breast carcinoma) | Exp Cell Res (2002) 281:50 | |
| Mitochondria | MitoTracker Red | M7512 | 578/599 | Sam5p (mitochondrial carrier for S-adenosylmethionine) | Yeast (Saccharomyces cerevisiae) | EMBO J (2003) 22:5975 | |
| Nuclear DNA | DAPI | D1306, D3571, D21490 | 358/461 | Histone H2B | HeLa | Methods (2003) 29:42 | |
| Nuclear DNA | Hoechst 33342 | H1399, H3570, H21492 | 350/461 | Histone H1 | BALB/c 3T3 fibroblasts | Nature (2000) 408:877 | |
| Nuclear DNA | SYTO 17 | S7579 | 621/634 | HIV-1 integrase | HeLa | J Biol Chem (2003) 278:33528 | |
| Nuclear DNA | SYTO 59 | S11341 | 622/645 | Microtubule plus-end binding protein | Porcine kidney epithelial cells (LLCPK) | Mol Biol Cell (2003) 14:916 | |
| Nuclear DNA | TO-PRO-3 | T3605 | 642/661 | Citron kinase | HeLa | J Cell Sci (2001) 114:3273 | |
| Plasma membrane | DiI | D282, D3911, N22880 | 549/565 | Synaptobrevin | Xenopus optic neurons | Nat Neurosci (2001) 4:1093 | |
| Other Subcellular Structures | |||||||
| F-actin | Rhodamine phalloidin | R415 | 554/573 | ERM (ezrin-radixin-moesin) proteins | Human peripheral blood T cells (PBT) | Nat Immunol (2004) 5:272 | |
| F-actin | Alexa Fluor 568 phalloidin | A12380 | 578/603 | Calponin | NIH 3T3 | J Cell Sci (2000) 113:3725 | |
| Lipid rafts | Cholera toxin subunit B (recombinant), Alexa Fluor 594 conjugate | C22842 | 590/617 | Histocompatibility leukocyte antigen (HLA)-Cw4 | NK cell–B-cell immunological synapse | Proc Natl Acad Sci U S A (2001) 98:14547 | |
| Cell Classification Markers | |||||||
| Apoptotic cells | Annexin V, Alexa Fluor 594 conjugate | A13203 | 590/617 | GRASP65 (Golgi stacking protein) | HeLa | J Cell Biol (2002) 156:495 | |
| Transformed B lymphocytes (Raji cells) | CellTracker Orange CMTMR | C2927 | 550/575 | ICAM-3 (intercellular adhesion molecule-3) | T-lymphocytes and antigen-presenting B cells | Nat Immunol (2002) 3:159 | |
| Cell-surface antigens | R-Phycoerythrin (streptavidin conjugate) | S866, S21388 | 565/575 | GFP gene expression | NIH 3T3 | Cytometry (1996) 25:211 | |
| Neurons | NeuroTrace 530/615 red-fluorescent Nissl stain | N21482 | 530/620 | Tau microtubule-binding protein (Purkinje cell marker) | Mouse brain slice | J Neurosci (2003) 23:6392 | |
| Neurons | Alexa Fluor 594 hydrazide | A10438, A10442 | 588/613 | Synaptophysin | Aplysia californica sensory neurons | Neuron (2003) 40:151 | |
| * This list covers only Aequoria victoria GFP, optimized mutants (e.g., EGFP) and green-fluorescent proteins from other species (e.g., Renilla reniformis). Fluorescent proteins with distinctly different excitation and emission characteristics (CFP, YFP, dsRed, etc.) are not included. † Fluorescence excitation (Ex) and emission (Em) maxima, in nm. ‡ Simultaneous imaging of GFP with fura-2 or ER-Tracker Blue-White DPX requires excitation wavelength–switching capability, because the fluorescence emission spectra overlap extensively. Even under these conditions, signal bleedthrough from one detection channel to the other may still be problematic, depending on the expression level and localization of the GFP chimera. See Biochem J (2001) 356:345 for further discussion. § The fluorescence emission spectra of styryl dyes such as FM 1-43 and FM 4-64 are broad and extend into the green emission range of GFP. In some cases, FM dye emission can overspill into the GFP detection channel, causing degraded resolution of image features. The excitation and emission spectra of FM 1-43 overlap those of GFP more extensively than those of FM 4-64. Therefore, using FM 4-64 instead of FM 1-43 is recommended to minimize this problem. | |||||||
Table 2. R0 values for FRET from EGFP to Alexa Fluor dyes.
| Acceptor Dye | R0 (Å)* |
|---|---|
| Alexa Fluor 546 dye | 57 |
| Alexa Fluor 555 dye | 63 |
| Alexa Fluor 568 dye | 54 |
| Alexa Fluor 594 dye | 53 |
| * R0 values in angstroms (Å) represent the distance at which fluorescence resonance energy transfer from the donor dye to the acceptor dye is 50% efficient. Values were calculated from spectroscopic data as outlined (Fluorescence Resonance Energy Transfer (FRET)—Note 1.2 ). |
For Research Use Only. Not for use in diagnostic procedures.