Search
Page Contents
- Quant-iT and Qubit Assay Kits for DNA and RNA
- Qubit 2.0 Fluorometer
- PicoGreen dsDNA Quantitation Assay
- OliGreen ssDNA Quantitation Assay
- RiboGreen RNA Quantitation Assay
- Other Stains for Nucleic Acid Quantitation in Solution
- Real-Time Quantitative PCR Using SYBR Green I Nucleic Acid Gel Stain
- Ordering Information
Related Tables
- Specialty nucleic acid reagents for molecular biology—Table 8.1
- Cell membrane–impermeant cyanine nucleic acid stains—Table 8.2
- Cell-permeant cyanine nucleic acid stains—Table 8.3
- Properties of classic nucleic acid stains—Table 8.4
- Characteristics of Molecular Probes ChromaTide and aha labeled nucleotides—Table 8.5
- Molecular Probes nucleic acid labeling kits—Table 8.6
- Amine-reactive dyes for nucleic acid sequencing—Table 8.7
- Selection guide for the Qubit and Quant-iT Assay Kits—Table 8.8
Related Technical Notes
Through intensive research efforts in both chemical synthesis and bioassay development, we have developed rapid and exceptionally sensitive fluorescence-based assays for quantitation of nucleic acids in solution. The Invitrogen Quant-iT Assay Kits represent advanced quantitation systems for DNA, RNA or protein samples. These state-of-the-art assays deliver high sensitivity and specificity together with a streamlined protocol, prediluted standards and a ready-to-use buffer. The Invitrogen Quant-iT PicoGreen dsDNA reagent, Quant-iT OliGreen ssDNA reagent and Quant-iT RiboGreen RNA reagent—optimized for double-stranded DNA, oligonucleotides and RNA, respectively—have a high affinity for nucleic acids and an extremely large fluorescence enhancement upon binding, making possible the direct detection of minute amounts of nucleic acids in complex solutions within minutes, usually without interference from other biomolecules. These reagents and quantitation assays provide the following advantages:
- Sensitivity. The Invitrogen PicoGreen dye–, OliGreen dye– and RiboGreen dye–based fluorescence assays are up to 10,000-fold more sensitive than UV absorbance measurements
and at least 400-fold more sensitive than assays that use the Hoechst 33258 dye
(H1398, H3569, H21491), requiring much less sample for quantitation.
- Accuracy. Unlike measurements of UV absorbance, these assays are not affected by the presence of proteins, free nucleotides or very short oligonucleotides, making quantitation of intact oligonucleotides and nucleic acids much more accurate in complex mixtures such as serum or whole blood.
- Precision. The average standard deviations of triplicate assays using these reagents are typically less than 5%.
- Simplicity. These assays have a very simple protocol that requires no separation steps, making them ideal for automated, high-throughput measurements.
- Broad dynamic range. Quantitation is accurate over four orders of magnitude for the PicoGreen and OliGreen assays, with a single dye concentration. The RiboGreen assay is accurate over three orders of magnitude.
- Instrument compatibility. Quantitation assays can be performed using a fluorescence microplate reader with standard filters optimized for fluorescein-like dyes, a relatively inexpensive filter-based spectrofluorometer or a standard spectrofluorometer. Additionally, we offer a set of Quant-iT Assay Kits specifically designed for the Invitrogen Qubit fluorometer.
- Convenience. Each reagent is available separately or in a kit that additionally contains a nuclease-free buffer and standards. The RiboGreen RNA quantitation assay is also available in the Invitrogen RediPlate 96 format—a prepared 96-well microplate with standards included—for high-throughput applications.
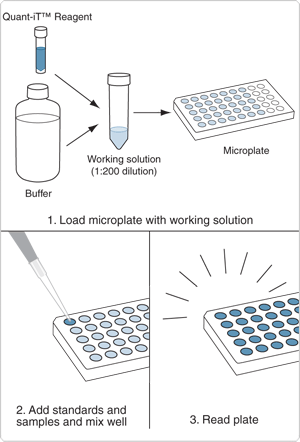
The Quant-iT family of assay kits—and the related Invitrogen Qubit family of assay kits designed for use with the Invitrogen Qubit 2.0 fluorometer—provides state-of-the-art reagents for sensitive and selective quantitation of DNA, RNA or protein samples using a standard fluorescence microplate reader or the Qubit 2.0 fluorometer (Selection Guide for the Qubit and Quant-iT Assay Kits—Table 8.8). These kits have been specially formulated with ready-to-use buffers, prediluted standards and easy-to-follow instructions, making quantitation both accurate and extremely easy (Figure 8.3.1). Each Quant-iT assay is:
- Ready to use. Only the dye is diluted in the supplied buffer; dilution of standards or buffer is not required.
- Easy to perform. Just add the sample to the diluted dye and read the fluorescence.
- Highly sensitive. The Quant-iT protein assay is orders of magnitude more sensitive than UV absorbance measurements.
- Highly selective. Separate kits are available for quantitating DNA, RNA (see below) or protein (Protein Quantitation in Solution—Section 9.2), with minimal interference from common contaminants.
- Precise. CVs are generally less than 5%.
Because the fluorescent dye in each Quant-iT Kit matches common fluorescence excitation and emission filter sets in microplate readers, these assay kits are ideal for high-throughput environments, as well as for small numbers of samples.
Quant-iT DNA Assay Kits
The Invitrogen Quant-iT DNA Assay Kits simplify DNA quantitation without sacrificing sensitivity. The Invitrogen Quant-iT dsDNA High-Sensitivity (HS) Assay Kit (Q33120) provides a linear detection range between 0.2 ng and 100 ng double-stranded DNA (dsDNA) (Figure 8.3.2), corresponding to initial experimental sample concentrations between 10 pg/µL and 100 ng/µL. This high-sensitivity DNA assay is ideal for quantitating PCR products, viral DNA, DNA fragments for subcloning and other applications requiring small amounts of DNA. The Invitrogen Quant-iT dsDNA Broad-Range (BR) Assay Kit (Q33130) provides a linear detection range between 2 ng and 1000 ng dsDNA (Figure 8.3.3), corresponding to initial experimental sample concentrations between 100 pg/µL and 1000 ng/µL. This broad-range DNA assay reduces the need to dilute concentrated samples, such as genomic DNA and miniprep DNA, prior to high-throughput procedures. Both Quant-iT DNA assays are highly selective for dsDNA over RNA, and have proven to be accurate even in the presence of an equal mass of RNA. Moreover, their fluorescence signals are unaffected by many common contaminants, including free nucleotides, salts, solvents and proteins. Each Quant-iT DNA Assay Kit contains:
- Invitrogen Quant-iT DNA HS reagent (in Kit Q33120) or Quant-iT DNA BR reagent (in Kit Q33130)
- Invitrogen Quant-iT DNA HS buffer (in Kit Q33120) or Quant-iT DNA BR buffer (in Kit Q33130)
- A set of eight prediluted λ DNA standards between 0 and 10 ng/µL (in Kit Q33120) or between 0 and 100 ng/µL (in Kit Q33130)
- Easy-to-follow instructions for the high-sensitivity DNA assay (Quant-iT dsDNA High-Sensitivity Assay Kit) or the broad-range DNA assay (Quant-iT dsDNA Broad-Range Assay Kit)
Sufficient reagents are provided to perform 1000 assays, based on a 200 µL assay volume in a 96-well microplate format; this assay can also be adapted for use in cuvettes or 384-well microplates. Both the high-sensitivity assay and the broad-range assay can be detected using standard fluorescein filters, and the fluorescence signal is stable for three hours at room temperature.
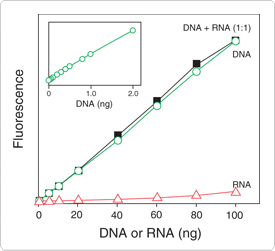
Figure 8.3.2 DNA selectivity and sensitivity of the Quant-iT dsDNA high-sensitivity assay. Triplicate 10 µL samples of DNA (![]() ), Escherichia coli rRNA (
), Escherichia coli rRNA (![]() ) or a 1:1 mixture of DNA and RNA (
) or a 1:1 mixture of DNA and RNA (![]() ) were assayed with the Quant-iT dsDNA High-Sensitivity Assay Kit (Q33120). Fluorescence was measured at 485/530 nm and plotted versus the mass of nucleic acid for the DNA alone, the mass of nucleic acid for the RNA alone or the mass of the DNA component in the 1:1 mixture. The coefficient of variation (CV) of replicate DNA determinations was ≤2%. The inset, a separate experiment with octuplicate determinations, shows the extreme sensitivity of the assay for DNA. Background fluorescence has not been subtracted.
) were assayed with the Quant-iT dsDNA High-Sensitivity Assay Kit (Q33120). Fluorescence was measured at 485/530 nm and plotted versus the mass of nucleic acid for the DNA alone, the mass of nucleic acid for the RNA alone or the mass of the DNA component in the 1:1 mixture. The coefficient of variation (CV) of replicate DNA determinations was ≤2%. The inset, a separate experiment with octuplicate determinations, shows the extreme sensitivity of the assay for DNA. Background fluorescence has not been subtracted.
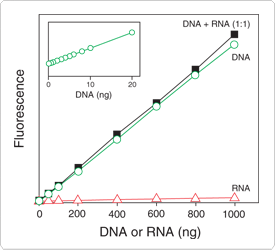
Figure 8.3.3 DNA selectivity and sensitivity of the Quant-iT dsDNA broad-range assay. Triplicate 10 µL samples of DNA (![]() ), Escherichia coli rRNA (
), Escherichia coli rRNA (![]() ) or a 1:1 mixture of DNA and RNA (
) or a 1:1 mixture of DNA and RNA (![]() ) were assayed with the Quant-iT dsDNA Broad-Range Assay Kit (Q33130). Fluorescence was measured at 485/530 nm and plotted versus the mass of nucleic acid for the DNA alone, the mass of nucleic acid for the RNA alone or the mass of the DNA component in the 1:1 mixture. The coefficient of variation (CV) of replicate DNA determinations was ≤3%. The inset, a separate experiment with octuplicate determinations, shows the sensitivity of the assay for DNA. Background fluorescence has not been subtracted.
) were assayed with the Quant-iT dsDNA Broad-Range Assay Kit (Q33130). Fluorescence was measured at 485/530 nm and plotted versus the mass of nucleic acid for the DNA alone, the mass of nucleic acid for the RNA alone or the mass of the DNA component in the 1:1 mixture. The coefficient of variation (CV) of replicate DNA determinations was ≤3%. The inset, a separate experiment with octuplicate determinations, shows the sensitivity of the assay for DNA. Background fluorescence has not been subtracted.
Quant-iT RNA Assay Kit
The Invitrogen Quant-iT RNA Assay Kit (Q33140) provides the first homogeneous assay ever developed for quantitating RNA in the presence of DNA. This RNA assay exhibits a linear detection range between 5 ng and 100 ng RNA (Figure 8.3.4), corresponding to initial experimental sample concentrations between 250 pg/µL and 100 ng/µL. We also offer the Invitrogen Quant-iT RNA Broad-Range Assay Kit (Q10213), with a linear detection range between 20 ng and 1000 ng. Because of the high selectivity of the Invitrogen Quant-iT RNA reagent for RNA over dsDNA, this assay enables accurate RNA quantitation even in the presence of an equal mass of DNA. The fluorescence signal is unaffected by many common contaminants, including free nucleotides, salts, solvents and proteins, making this assay ideal for measuring samples for microarray, RT-PCR and northern blot procedures. Each Quant-iT RNA Assay Kit contains:
- Quant-iT RNA reagent
- Invitrogen Quant-iT RNA buffer
- A set of eight prediluted Escherichia coli rRNA standards between 0 and 10 ng/µL
- Easy-to-follow instructions (Quant-iT RNA Assay Kit)
Sufficient reagents are provided to perform 1000 assays, based on a 200 µL assay volume in a 96-well microplate format; this assay can also be adapted for use in cuvettes or 384-well microplates. The fluorescence signal exhibits excitation/emission maxima of 644/673 nm and is stable for three hours at room temperature.
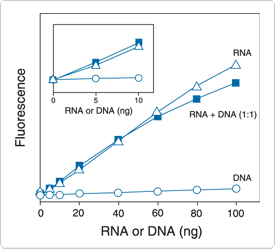
Figure 8.3.4 RNA selectivity and sensitivity of the Quant-iT RNA assay. Triplicate 10 µL samples of Escherichia coli rRNA (![]() ), DNA (
), DNA (![]() ) or a 1:1 mixture of RNA and DNA (
) or a 1:1 mixture of RNA and DNA (![]() ) were assayed with the Quant-iT RNA Assay Kit (Q33140). Fluorescence was measured at 630/680 nm and plotted versus the mass of nucleic acid for the RNA alone, the mass of nucleic acid for the DNA alone or the mass of the RNA component in the 1:1 mixture. The coefficient of variation (CV) of replicate RNA determinations was ≤10%. The inset is an enlargement of the graph to show the sensitivity of the assay for RNA. Background fluorescence has not been subtracted.
) were assayed with the Quant-iT RNA Assay Kit (Q33140). Fluorescence was measured at 630/680 nm and plotted versus the mass of nucleic acid for the RNA alone, the mass of nucleic acid for the DNA alone or the mass of the RNA component in the 1:1 mixture. The coefficient of variation (CV) of replicate RNA determinations was ≤10%. The inset is an enlargement of the graph to show the sensitivity of the assay for RNA. Background fluorescence has not been subtracted.
Qubit DNA and RNA Assay Kits for Use with the Qubit Fluorometer
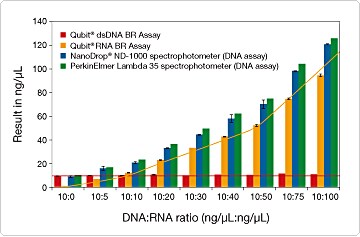
In addition to the Quant-iT Assay Kits for microplate readers (described above), we offer a set of Qubit Assay Kits specifically formulated for use with the Qubit 2.0 fluorometer (Selection Guide for the Qubit and Quant-iT Assay Kits—Table 8.8); these kits are also compatible with the original Invitrogen Qubit (1.0) fluorometer. As with the Quant-iT DNA and RNA assays, the most significant advantage of the Invitrogen Qubit DNA and RNA assays is their selectivity. UV absorbance–based analysis cannot be used to quantitate DNA or RNA in samples containing a mixture of both. In contrast, the Qubit DNA and RNA assays are able to accurately measure DNA and RNA, respectively, in the same sample (Figure 8.3.5). We found that the DNA concentration of a sample containing equal parts DNA and RNA can be measured within 2% of the actual concentration using the Invitrogen Qubit dsDNA BR (Broad-Range) Assay Kit. Furthermore, in a sample containing a 10-fold excess of RNA over DNA, the DNA concentration was measured to be only 7% higher than the actual concentration using the Qubit dsDNA BR Assay Kit.
We offer a variety of Invitrogen Qubit DNA and RNA Assay Kits that differ in their selectivity and sensitivity range:
Each Qubit Assay Kit provides concentrated assay reagent, dilution buffer and prediluted standards; all assay reagents are stored at room temperature. The assay protocol is fast and easy: Simply dilute the reagent using the buffer provided, add the sample (any volume between 1 and 20 μL) and, after a 2-minute incubation, read the concentration using the Qubit fluorometer. With the Qubit 2.0 fluorometer, all assay settings and calculations are performed automatically, and the Qubit assay signal is stable for 3 hours.
Accurate and sensitive nucleic acid and protein quantitation demands the highest levels of performance, from the analyte-specific assay straight through to the analytical instrument. The Qubit 2.0 fluorometer (Figure 8.3.6) is our second-generation desktop fluorometer designed to work seamlessly with the Qubit DNA, RNA and protein assays (formerly called Quant-iT assays). Together, they form the Invitrogen Qubit Fluorometric Quantitation Platform, an efficient combination of sophisticated, accurate, and highly sensitive fluorescence-based quantitation assays, along with an extremely user-friendly fluorometer. This powerful pairing offers:
- Selective quantitation—more selective and accurate than UV absorbance readings
- High sensitivity—use as little as 1 µL of sample for quantitation
- Intuitive integrated platform—sophisticated quantitation in 5 minutes or less
The Qubit 2.0 fluorometer offers state-of-the art optical components, sophisticated data analysis algorithms and a small bench footprint. It provides the same accuracy provided by the original Qubit (1.0) fluorometer, but with even more speed and less effort than before. The Qubit 2.0 fluorometer has an upgraded, robust design, complete with a color touch screen and intuitive user interface. Navigation buttons for running samples, calibrating standards, and reviewing results are continually accessible at the bottom of the screen, eliminating multiple steps and confusion. And no more scrolling through selections one-by-one using “Up” or “Down” arrows—the Qubit 2.0 fluorometer is equipped with list wheels and scroll bars for clear and simple line selection. Importantly, all functionality has been validated for convenient use with gloves, such as latex and nitrile, commonly worn in a lab setting.
Additionally, the Qubit 2.0 fluorometer includes a new graphing feature that shows the standard curve after calibration and displays up to 10 data points. Sample data is automatically stored in a .csv file format and can be accessed at any time by navigating to the data screen. This stored data file contains all relevant sample information, including time/date stamp, concentration readout, units, and assay type, as well as raw fluorescence data for assay troubleshooting. A built-in dilution calculator allows users to quickly determine sample content, and concentrations determined using this calculator are also stored in the data file. Furthermore, although each Qubit assay function comes pre-programmed with default unit readouts, the Qubit 2.0 fluorometer enables users to change unit output based on their own preference.
For simpler identification and sorting, the data files can be renamed directly using the instrument‘s full alphanumeric keypad. This data can then be transferred to a computer through a USB drive for more thorough analysis—no direct connection to a PC is required at anytime, neither for instrument operation nor data transfer. Likewise, any upgrades can easily be made using a USB drive and firmware downloaded from www.invitrogen.com/qubit. For convenience, every Qubit 2.0 fluorometer comes with a 2 GB USB drive equipped with .pdf files of the complete instrument user manual and all Qubit Assay Kit protocols. We offer the Qubit fluorometer with USB drive separately (Q32866; replacement USB drive, Q32867; replacement international power cord, Q32868) or bundled in a starter kit that includes four different Qubit assay kits as well as a supply of Qubit assay tubes (Q32871). A set of 500 Qubit assay tubes is also available separately (Q32856).
Qubit Assay Kits use advanced Molecular Probes fluorophores that become fluorescent upon binding to DNA, RNA or protein. The selectivity of these interactions ensures more accurate results than can be obtained with UV absorbance readings; Qubit Assay Kits only report the concentration of the molecule of interest, not sample contaminants. Qubit Assay Kits are up to 1000 times as sensitive as UV absorbance readings, and as little as 1 µL of sample is all that's needed for accurate, reliable quantitation. We offer Qubit Assay Kits for a variety of quantitation needs:
- Qubit dsDNA Broad-Range (BR) and High-Sensitivity (HS) Assay Kits—for sequencing samples, genomic DNA samples and routine cloning experiments (see above)
- Qubit RNA Assay Kits—for microarray experiments, real-time PCR samples and northern blot analysis (see above)
- Qubit Protein Assay Kits—for protein samples destined for SDS-PAGE, 2D gel electrophoresis and western blot analysis (Q33211, Q33212; Protein Quantitation in Solution—Section 9.2)

Figure 8.3.6 The Qubit 2.0 fluorometer.
The Quant-iT PicoGreen dsDNA reagent ![]() (P7581, P11495) and Kits (P7589, P11496) can accurately quantitate as little as 25 pg/mL of dsDNA in a fluorometer or 250 pg/mL (typically 50 pg in a 200 µL volume) in a fluorescence microplate reader. The PicoGreen dsDNA quantitation assay is more than 10,000 times as sensitive as conventional UV absorbance measurements at 260 nm
(P7581, P11495) and Kits (P7589, P11496) can accurately quantitate as little as 25 pg/mL of dsDNA in a fluorometer or 250 pg/mL (typically 50 pg in a 200 µL volume) in a fluorescence microplate reader. The PicoGreen dsDNA quantitation assay is more than 10,000 times as sensitive as conventional UV absorbance measurements at 260 nm ![]() (an A260 of 0.1 corresponds to an ~5 µg/mL dsDNA solution) and at least 400 times more sensitive than the Hoechst 33258 dye–based assay.
(an A260 of 0.1 corresponds to an ~5 µg/mL dsDNA solution) and at least 400 times more sensitive than the Hoechst 33258 dye–based assay.![]() It is even more sensitive than assays based on our YO-PRO-1 and YOYO-1 dyes, which have reported detection limits of approximately 2.5 ng/mL and 0.5 ng/mL,
It is even more sensitive than assays based on our YO-PRO-1 and YOYO-1 dyes, which have reported detection limits of approximately 2.5 ng/mL and 0.5 ng/mL,![]() respectively.
respectively.
Although the PicoGreen reagent is not specific for dsDNA, it shows a >1000-fold fluorescence enhancement upon binding to dsDNA, and much less fluorescence enhancement upon binding to single-stranded DNA (ssDNA) or RNA, making it possible to quantitate dsDNA in the presence of equimolar amounts of ssDNA, RNA or proteins ![]() (Figure 8.3.7). The PicoGreen reagent also selectively detects DNA–RNA hybrids in the presence of ssDNA and RNA. Differences in the emission lifetimes of the PicoGreen complexes with dsDNA and ssDNA make it possible to quantitate the relative amounts of each species in solution by time-resolved measurements. By contrast, UV absorbance measurements cannot distinguish between dsDNA, ssDNA and RNA or proteins. Thus, the PicoGreen reagent allows direct quantitation of PCR amplicons without purification from the reaction mixture and makes it possible to detect low levels of DNA contamination in recombinant protein products.
(Figure 8.3.7). The PicoGreen reagent also selectively detects DNA–RNA hybrids in the presence of ssDNA and RNA. Differences in the emission lifetimes of the PicoGreen complexes with dsDNA and ssDNA make it possible to quantitate the relative amounts of each species in solution by time-resolved measurements. By contrast, UV absorbance measurements cannot distinguish between dsDNA, ssDNA and RNA or proteins. Thus, the PicoGreen reagent allows direct quantitation of PCR amplicons without purification from the reaction mixture and makes it possible to detect low levels of DNA contamination in recombinant protein products.![]() In comparison to the Hoechst 33258 dye, which shows significant AT selectivity, the PicoGreen reagent shows little if any AT- or GC-selectivity, enabling accurate DNA quantitation from many sources.
In comparison to the Hoechst 33258 dye, which shows significant AT selectivity, the PicoGreen reagent shows little if any AT- or GC-selectivity, enabling accurate DNA quantitation from many sources.
The protocol for the PicoGreen dsDNA quantitation assay is simple and requires very few steps—the dye is simply added to the sample and incubated for five minutes, then the fluorescence is measured. In addition, the fluorescence signal from binding of the PicoGreen reagent to dsDNA is linear over at least four orders of magnitude (Figure 8.3.8) with a single dye concentration, whereas assays using ethidium bromide, Hoechst 33258 or the YOYO-1 dye exhibit a much more limited linear range.![]() We have found that this linearity is maintained in the presence of several compounds commonly found in nucleic acid preparations, including salts, urea, ethanol, chloroform, detergents, proteins and agarose.
We have found that this linearity is maintained in the presence of several compounds commonly found in nucleic acid preparations, including salts, urea, ethanol, chloroform, detergents, proteins and agarose.![]() The PicoGreen reagent can be excited at 488 nm with an argon-ion laser, and is reported to be a superior nucleic acid stain for analysis of single DNA molecules in a flow cytometer.
The PicoGreen reagent can be excited at 488 nm with an argon-ion laser, and is reported to be a superior nucleic acid stain for analysis of single DNA molecules in a flow cytometer.![]() A method that utilizes the PicoGreen reagent for the absolute quantitation of cDNA using real-time PCR has been reported.
A method that utilizes the PicoGreen reagent for the absolute quantitation of cDNA using real-time PCR has been reported.![]()
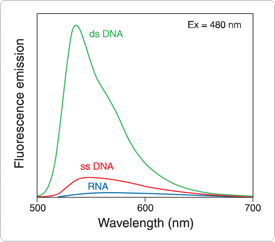
Figure 8.3.7. Fluorescence enhancement of the Quant-iT PicoGreen dsDNA reagent upon to binding dsDNA, ssDNA and RNA. Samples containing 500 ng/mL calf thymus DNA, M13 ssDNA or Escherichia coli ribosomal RNA were added to cuvettes containing Quant-iT PicoGreen dsDNA reagent (P7581, P7589, P11495, P11496) in TE buffer. Samples were excited at 480 nm, and the fluorescence emission spectra were collected using a spectrofluorometer. Emission spectra for samples containing dye and nucleic acids, as well as for dye alone (baseline), are shown.
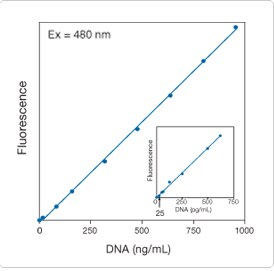
Figure 8.3.8. Linear quantitation of calf thymus DNA from 25 pg/mL to 1000 ng/mL using the Quant-iT PicoGreen dsDNA reagent (P7581, P7589, P11495, P11496). Samples in 10 mm × 10 mm cuvettes were excited at 480 nm. The fluorescence emission intensity was measured at 520 nm using a spectrofluorometer and plotted as a function of DNA concentration. The inset shows an enlargement of the results obtained with DNA concentrations between 0 and 750 pg/mL.
The PicoGreen assay is useful for quantitating DNA templates for PCR,![]() labeling reactions, electrophoretic mobility-shift (bandshift) assays, DNA-footprinting assays and filter-binding assays, and for measuring yields from PCR reactions,
labeling reactions, electrophoretic mobility-shift (bandshift) assays, DNA-footprinting assays and filter-binding assays, and for measuring yields from PCR reactions,![]() DNA minipreps and maxipreps, cDNA synthesis and nuclease protection assays. The simplicity and selectivity of the assay also make it ideal for high-throughput automated quantitation assays used in forensic and genomics research. Furthermore, the PicoGreen reagent has been used for:
DNA minipreps and maxipreps, cDNA synthesis and nuclease protection assays. The simplicity and selectivity of the assay also make it ideal for high-throughput automated quantitation assays used in forensic and genomics research. Furthermore, the PicoGreen reagent has been used for:
- Genotyping by allele-specific PCR

- Quantitating dsDNA samples before and after PCR amplification or after agarose gel electrophoresis

- Determining PCR amplification yields before sequencing

- Automating quantitation of DNA isolated from biological samples or obtained from PCR reactions, prior to running DNA typing gels for high-throughput genotyping

- Quantitating DNA from buccal scrapes prior to DNA profiling by short tandem repeat (STR) analysis

- Identifying contaminating DNA in recombinant protein products
 or purified monoclonal antibody preparations
or purified monoclonal antibody preparations 
- Monitoring DNA strand breaks in plasmids and bacterial artificial chromosomes

- Quantitating the efficiency of DNA extraction from frozen and formalin-fixed tissue sections

- Detecting mammalian telomerase activity in tumor cells using the PCR-based TRAP assay

- Developing assays for DNA polymerase and reverse transcriptase

- Measuring the activity of DNase I

- Measuring supercoiled DNA forms in solution, based on their renaturation properties

- Monitoring plasmid production during fermentation and downstream processing

- Quantitating DNA denaturation as a measure for DNA damage in purified DNA preparations, cell lysates or homogenized solid tissues

Each vial of the Quant-iT PicoGreen dsDNA reagent (P7581) contains a sufficient amount of dye for at least 200 assays using a 2 mL assay volume and a standard fluorometer, or 2000 assays using a 200 µL assay volume and a fluorescence microplate reader. The product is accompanied by a simple protocol that enables linear and reproducible quantitation of dsDNA. We also provide the PicoGreen reagent in the Quant-iT PicoGreen dsDNA Assay Kit (P7589), which contains:
- PicoGreen dsDNA quantitation reagent
- Low-fluorescence, nucleic acid– and nuclease-free assay buffer concentrate, essential for the accurate measurement of dsDNA
- dsDNA standard solution for assay calibration
- Detailed protocols for dsDNA quantitation (Quant-iT PicoGreen dsDNA Reagent and Kits)
This kit provides sufficient reagents for 200 assays using a 2 mL assay volume and a standard fluorometer or 2000 assays using a 200 µL assay volume and a fluorescence microplate reader. Both the stand-alone reagent and the kit are also available in special packaging, in which the PicoGreen reagent is supplied as 10 vials of 100 µL aliquots for added convenience (P11495; Kit, P11496). The special packaging reduces thawing times, provides individual aliquots for each person performing the assay, and allows smaller amounts of dye to be taken into the field for analysis of water or other samples.
For researchers working with ssDNA and oligonucleotides, as well as for companies that synthesize oligonucleotides, we offer the Quant-iT OliGreen ssDNA reagent (O7582) and Quant-iT OliGreen ssDNA Assay Kit (O11492). Short, synthetic oligonucleotides are used in a number of molecular biology techniques, including DNA sequencing, site-directed mutagenesis, DNA amplification, antisense gene suppression and in situ and blot hybridization. The conventional methods for quantitating oligonucleotides are not very sensitive, often requiring highly concentrated samples, and are quite subject to interference from sample contaminants. The most commonly used technique for measuring oligonucleotide and ssDNA concentrations is the determination of absorbance at 260 nm (an A260 of 0.1 corresponds to an ~3 µg/mL solution of a synthetic 24-mer M13 sequencing primer).
The Quant-iT OliGreen ssDNA reagent enables researchers to routinely quantitate as little as 100 pg/mL of ssDNA or oligonucleotide (200 pg in a 2 mL assay volume with a standard fluorometer) or 200 pg in a 200 µL assay volume using a fluorescence microplate reader (Figure 8.3.9). Thus, quantitation with the OliGreen reagent is about 10,000 times more sensitive than quantitation with UV absorbance methods and at least 500 times more sensitive (and far faster, with a greater throughput) than detecting oligonucleotides on electrophoretic gels stained with ethidium bromide. Using an easy-to-follow protocol and fluorescein excitation and emission wavelengths, we have quantitated oligonucleotides that range from 10 to 50 nucleotides in length, as well as several sources of ssDNA, such as M13 and φX174 viral DNA and denatured calf thymus DNA, and obtained similar sensitivity. The OliGreen reagent has also been used to detect phosphodiester and phosphorothioate oligonucleotides.![]()
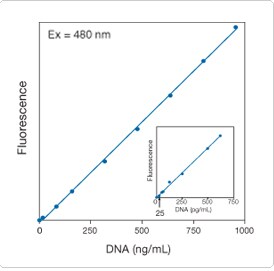
Figure 8.3.9. Linear quantitation of a synthetic 24-mer (an M13 sequencing primer) from 0.1 to 1000 ng/mL using the Quant-iT OliGreen ssDNA reagent (O7582, O11492). Samples in 10 mm × 10 mm cuvettes were excited at 480 nm. The fluorescence emission intensity was measured at 520 nm using a spectrofluorometer and plotted as a function of oligonucleotide concentration. The inset shows an enlargement of the results obtained with oligonucleotide concentrations between zero and 2.0 ng/mL.
Significant disadvantages of the UV absorbance method for oligonucleotide quantitation include the large relative contribution of free nucleotides to the signal and the interference caused by contaminants commonly found in nucleic acid preparations. By contrast, nucleotides and short oligonucleotides of six bases or less do not interfere with the OliGreen ssDNA quantitation assay. However, the OliGreen reagent does exhibit fluorescence enhancement when bound to dsDNA and RNA. Like the PicoGreen assay, the linear detection range of the OliGreen ssDNA quantitation assay in a standard fluorometer extends over four orders of magnitude—from 100 pg/mL to 1 µg/mL—with a single dye concentration (Figure 8.3.9). The linearity of the OliGreen assay is maintained in the presence of several compounds commonly found to contaminate nucleic acid preparations, including salts, urea, ethanol, chloroform, detergents, proteins, ATP and agarose; however, many of these compounds do affect the signal intensity to some extent, so standard curves should be generated using solutions that closely mimic those of the samples. The OliGreen assay can even be performed using samples as complex as whole blood or serum.![]()
Our experiments with homopolymers have demonstrated that the OliGreen reagent may exhibit significant base selectivity. The OliGreen reagent shows a large fluorescence enhancement when bound to poly(dT) but only a relatively small fluorescence enhancement when bound to poly(dG) and little signal with poly(dA) and poly(dC). Thus, it is important to use an oligonucleotide with similar base composition when generating the standard curve.
The remarkable properties of our OliGreen reagent make it ideal for the fast and accurate detection and quantitation of:
Each vial of Quant-iT OliGreen ssDNA reagent (O7582) contains a sufficient amount of dye for at least 200 assays using a 2 mL assay volume and a standard fluorometer, or 2000 assays using a 200 µL assay volume and a fluorescence microplate reader. The product is accompanied by a simple protocol that ensures linear and reproducible quantitation of ssDNA. We also provide the OliGreen reagent in the Quant-iT OliGreen ssDNA Assay Kit (O11492), which contains:
- OliGreen ssDNA quantitation reagent
- Low-fluorescence, nucleic acid– and nuclease-free assay buffer concentrate, essential for the accurate measurement of ssDNA
- Oligonucleotide standard (M13 sequencing primer) solution for assay calibration
- Detailed protocols for ssDNA quantitation (Quant-iT OliGreen ssDNA Reagent and Kit)
This kit provides sufficient reagents for 200 assays using a 2 mL assay volume and a standard fluorometer or 2000 assays using a 200 µL assay volume and a fluorescence microplate reader.
RiboGreen RNA Reagent and Kit
The Quant-iT RiboGreen RNA reagent (R11491) is our premier stain for quantitating RNA in solution. Like the PicoGreen dsDNA and OliGreen ssDNA quantitation assays, the RiboGreen RNA quantitation assay relies on a proprietary dye that exhibits a large fluorescence enhancement upon binding to nucleic acids. The extinction coefficient (EC) of the RiboGreen reagent, as well as its quantum yield (QY) and fluorescence enhancement upon binding RNA, are all significantly greater than those of ethidium bromide.
- RiboGreen reagent: EC482 nm = 67,000 cm-1M-1, QY = 0.65, fluorescence enhancement >1000-fold.
- Ethidium bromide: EC482 nm = 5,500 cm-1M-1, QY <0.3, fluorescence enhancement <30-fold.
The RiboGreen assay allows detection of as little as 1 ng/mL RNA in a standard fluorometer, fluorescence microplate reader or filter-based fluorometer using standard fluorescein excitation and emission settings (Figure 8.3.10). This sensitivity is at least 200-fold better than that achieved with ethidium bromide ![]() and at least 1000-fold better than that achieved using conventional absorbance measurements at 260 nm (an A260 of 0.1 corresponds to an ~4 µg/mL RNA solution). Unlike UV absorbance measurements at 260 nm, the RiboGreen reagent does not detect significant sample contamination by free nucleotides.
and at least 1000-fold better than that achieved using conventional absorbance measurements at 260 nm (an A260 of 0.1 corresponds to an ~4 µg/mL RNA solution). Unlike UV absorbance measurements at 260 nm, the RiboGreen reagent does not detect significant sample contamination by free nucleotides.![]() Thus, the RiboGreen reagent more accurately measures the amount of intact RNA polymers in potentially degraded samples.
Thus, the RiboGreen reagent more accurately measures the amount of intact RNA polymers in potentially degraded samples.
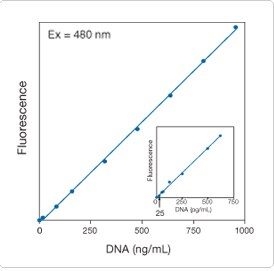
Figure 8.3.10. Linear quantitation of ribosomal RNA using the Quant-iT RiboGreen RNA reagent (R11491, R11490). For the high-range assay, the RiboGreen reagent was diluted 200-fold into 10 mM Tris-HCl, 1 mM EDTA, pH 7.5 (TE), and 100 µL of the reagent solution was added to microplate wells containing 100 µL ribosomal RNA in TE. For the low-range assay (see inset), the RiboGreen reagent was diluted 2000-fold into TE, and 100 µL of the reagent solution was added to 100 µL of ribosomal RNA in TE. Samples were excited at 485 ± 10 nm, and the fluorescence emission intensity was measured at 530 ± 12.5 nm using a fluorescence microplate reader. Background fluorescence was not subtracted.
Using two different dye concentrations to cover its full dynamic range of three orders of magnitude, we have observed a linear correlation between the RNA concentration and fluorescence for 1.0 ng/mL to 50 ng/mL RNA using a 4000-fold dilution of the RiboGreen reagent, and for 20 ng/mL to 1 µg/mL using a 400-fold dilution of the dye (Figure 8.3.10). Assay linearity is maintained in the presence of several compounds commonly found in nucleic acid preparations, including salts, urea, ethanol, chloroform, detergents, proteins and agarose.![]()
The RiboGreen reagent is not appreciably selective for RNA—the dye also shows significant fluorescence enhancement upon binding to DNA. However, a simple DNase pretreatment of samples removes the contribution of DNA to the signal. The RiboGreen reagent may also have some base selectivity; it exhibits about 60% less fluorescence when bound to poly(G) homopolymers and virtually no fluorescence when bound to poly(U) or poly(C) homopolymers compared with the fluorescence when bound to poly(A) homopolymers or to rRNA.![]()
Using the Quant-iT RiboGreen RNA reagent, we have reproducibly quantitated RNA from a wide variety of sources, including ribosomal RNA (rRNA), transfer RNA (tRNA), viral RNA, polyA+ fractions and total cellular RNA.![]() The RiboGreen reagent is useful for:
The RiboGreen reagent is useful for:
- Fast and accurate measurements of RNA yields before generating cDNA
- Determination of RNA yields from in vitro transcription
- Accurate measurements of RNA before performing northern blot analysis,
S1 nuclease assays, RNase-protection assays, reverse-transcription PCR and differential-display PCR
- Assay of DNA-dependent RNA polymerase activity
- Detection of capillary electrophoresis–separated viral RNA that has been stained in vitro
Each vial of the Quant-iT RiboGreen RNA reagent (R11491) contains a sufficient amount of dye for at least 200 high-range assays or 2000 low-range assays using a 2 mL assay volume and a standard fluorometer. With a fluorescence microplate reader and a 96-well microplate, the assay volume is reduced to 200 µL, allowing 2000 high-range assays or 20,000 low-range assays. Included with each vial of the RiboGreen reagent is a simple protocol that enables linear and reproducible quantitation of RNA. We also provide the RiboGreen reagent in the Quant-iT RiboGreen RNA Assay Kit (R11490), which contains:
- RiboGreen RNA reagent
- Low-fluorescence, nucleic acid– and nuclease-free assay buffer concentrate, essential for the accurate measurement of RNA
- Ribosomal RNA standard (16S and 23S rRNA from Escherichia coli) solution for assay calibration
- Detailed protocols (Quant-iT RiboGreen RNA Reagent and Kit)
This kit provides sufficient reagents for at least 200 assays using a 2 mL assay volume and a standard fluorometer or at least 2000 assays using a 200 µL assay volume and a fluorescence microplate reader. The RNase-free TE buffer concentrate, which is essential to the success of the assay, is also available separately (T11493) and can be used to extend the number of low-concentration assays possible with the kit.
RediPlate 96 RiboGreen RNA Quantitation Kit
The RiboGreen assay is also available in a convenient Invitrogen RediPlate 96 RiboGreen RNA Quantitation Kit in which the RiboGreen reagent is predispensed into a 96-well microplate (R32700). The buffer and sample are simply added to the microplate wells—there is no need to handle the RiboGreen reagent. After a 10-minute incubation, the microplate is ready to read in a fluorescence microplate reader. The RediPlate RiboGreen assay has a linear range of ~15–1000 ng/mL (~3–200 ng in a 200 µL assay volume) with a single dye concentration (Figure 8.3.11). The microplate used in the RediPlate 96 RiboGreen RNA Quantitation Kit is provided in a resealable foil packet, and it snaps apart into twelve strips to permit assays in any multiple of eight (Figure 8.3.12). Eleven of the strips are preloaded with the RiboGreen reagent; the remaining strip, marked with blackened tabs, contains a series of RNA standards for generating a standard curve. In addition to the 96-well microplate, each RediPlate RiboGreen 96 RNA Quantitation Kit includes RNase-free reaction buffer and detailed instructions (RediPlate 96 RiboGreen RNA Quantitation Kit).
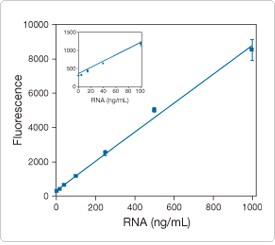
Figure 8.3.11. Dynamic range and sensitivity of the RediPlate 96 RiboGreen RNA quantitation assay. The RNA standards provided in the RediPlate 96 RiboGreen RNA Quantitation Kit (R32700) were added in quadruplicate to assay wells as described in the accompanying protocol, and fluorescence was measured in a fluorescence microplate reader using excitation at 485 ± 12.5 nm and fluorescence detection at 530 ± 15 nm. Fluorescence was plotted against the RNA concentration with no background subtraction. The inset shows the sensitivity of the assay at very low levels of RNA.

Figure 8.3.12. A RediPlate 96 microplate.
Cyanine Dyes and Phenanthridine Dyes for Nucleic Acid Quantitation in Solution
The dimeric cyanine dyes Invitrogen TOTO-1 and YOYO-1 are useful for sensitive fluorometric measurement of dsDNA, ssDNA and RNA in solution,![]() although the PicoGreen, OliGreen and RiboGreen reagents generally are faster, have greater sensitivity and show a linear response over a broader range of nucleic acid concentrations. The linear range of assays that use the TOTO-1 and YOYO-1 dyes for DNA quantitation encompasses about two orders of magnitude, with a sensitivity limit of about 0.5 ng/mL. The Invitrogen TOTO-1, YOYO-1 and YO-PRO-1 nucleic acid stains have been used to quantitate PCR amplification products in a homogeneous human leukocyte antigen (HLA) typing method that requires no transfer or washing steps, thus minimizing the risk of sample contamination.
although the PicoGreen, OliGreen and RiboGreen reagents generally are faster, have greater sensitivity and show a linear response over a broader range of nucleic acid concentrations. The linear range of assays that use the TOTO-1 and YOYO-1 dyes for DNA quantitation encompasses about two orders of magnitude, with a sensitivity limit of about 0.5 ng/mL. The Invitrogen TOTO-1, YOYO-1 and YO-PRO-1 nucleic acid stains have been used to quantitate PCR amplification products in a homogeneous human leukocyte antigen (HLA) typing method that requires no transfer or washing steps, thus minimizing the risk of sample contamination.![]() Other dyes for nucleic acid quantitation in solution and their applications include:
Other dyes for nucleic acid quantitation in solution and their applications include:
- YOYO-1 dye (Y3601, Nucleic Acid Stains—Section 8.1) for solution quantitation of oligonucleotides,
PCR products
and nuclear run-on assays
- YO-PRO-1 dye (Y3603, Nucleic Acid Stains—Section 8.1) for quantitating dsDNA in a fluorescence microplate reader, with a reported sensitivity limit of about 2.5 ng/mL
- YO-PRO-1 dye for quantitating RNA isolated from Xenopus embryos
- YO-PRO-1 dye and Invitrogen SYBR Green I dye for direct counting of viruses in marine and freshwater environments
(
)
- Invitrogen PO-PRO-3 dye (P3585, Nucleic Acid Stains—Section 8.1) for quantitating DNA in a fluorescence microplate reader,
with fluorescence measurements reported to be independent of base-pair composition
- Ethidium bromide for measuring the yield of PCR products
- Ethidium bromide and acridine homodimer (A666, Nucleic Acid Stains—Section 8.1) for quantitating covalently closed, circular DNA and for measuring the activity of polymerases, deoxynucleotidyl transferases, ligases, gyrases, topoisomerases and nucleases
- YOYO-1 dye for measuring the activity of DNases
- Invitrogen SYBR Gold nucleic acid gel stain (S11494, Nucleic Acid Detection on Gels, Blots and Arrays— Section 8.4), for detecting DNA mutations in a simple PCR-based assay
Dyes such as the ethidium homodimers and our dimeric cyanine dyes—the Invitrogen TOTO, YOYO, BOBO, POPO, JOJO and LOLO dyes (Cell membrane–impermeant cyanine nucleic acid stains—Table 8.2)—exhibit a high affinity for double-stranded nucleic acids but label small single-stranded oligonucleotides less well. This characteristic of ethidium homodimer-1 (E1169, Nucleic Acid Stains—Section 8.1) was exploited to analyze short self-annealing oligonucleotides for their ability to hybridize.![]() Because our dimeric cyanine dyes and the Invitrogen PicoGreen dsDNA quantitation reagent have extremely low intrinsic fluorescence in the absence of DNA, high fluorescence enhancements upon binding, higher quantum yields and much larger extinction coefficients than ethidium homodimer-1,
Because our dimeric cyanine dyes and the Invitrogen PicoGreen dsDNA quantitation reagent have extremely low intrinsic fluorescence in the absence of DNA, high fluorescence enhancements upon binding, higher quantum yields and much larger extinction coefficients than ethidium homodimer-1,![]() they should prove superior in this application.
they should prove superior in this application.
Hoechst 33258 Dye for Quantitating DNA in Solution
The Hoechst 33258 dye (H1398, H3569, H21491) has been extensively used to quantitate dsDNA in solution. Hoechst 33258 shows a fluorescence increase upon binding nucleic acids and a preference for binding to AT regions. Hoechst 33258 is selective (but not specific) for dsDNA over RNA in high-salt buffers and for dsDNA over ssDNA in low-salt buffers. The Hoechst 33258 dye can quantitatively detect from 10 ng/mL to ~10 µg/mL dsDNA when two different dye concentrations are used.![]() While this assay uses principles that are similar to other fluorescent assays, newer dyes such as the PicoGreen reagent provide much higher sensitivity, better selectivity and a much broader dynamic range with a single dye concentration.
While this assay uses principles that are similar to other fluorescent assays, newer dyes such as the PicoGreen reagent provide much higher sensitivity, better selectivity and a much broader dynamic range with a single dye concentration.
The Invitrogen FluoReporter Blue Fluorometric dsDNA Quantitation Kit (F2962) provides the protocols developed by Rago and colleagues ![]() for analyzing cellular DNA with the blue-fluorescent Hoechst 33258 nucleic acid stain. The kit enables researchers to detect ~10 ng of isolated calf thymus DNA or ~1000 mouse NIH 3T3 cells in a 200 µL sample (substantially lower levels are detectable using our Invitrogen CyQUANT Cell Proliferation Assay Kit described in Assays for Cell Enumeration, Cell Proliferation and Cell Cycle—Section 15.4). With this kit, quantitation of cellular DNA is rapid, and all manipulations can be carried out in microplate wells. The cells are lysed by freezing them in distilled water, which circumvents the requirement for extraction procedures used in other Hoechst 33258 dye–based protocols.
for analyzing cellular DNA with the blue-fluorescent Hoechst 33258 nucleic acid stain. The kit enables researchers to detect ~10 ng of isolated calf thymus DNA or ~1000 mouse NIH 3T3 cells in a 200 µL sample (substantially lower levels are detectable using our Invitrogen CyQUANT Cell Proliferation Assay Kit described in Assays for Cell Enumeration, Cell Proliferation and Cell Cycle—Section 15.4). With this kit, quantitation of cellular DNA is rapid, and all manipulations can be carried out in microplate wells. The cells are lysed by freezing them in distilled water, which circumvents the requirement for extraction procedures used in other Hoechst 33258 dye–based protocols.![]() The diluted dye solution is then added to lysed cells and the fluorescence is measured. Kit components include:
The diluted dye solution is then added to lysed cells and the fluorescence is measured. Kit components include:
- Hoechst 33258 in dimethylsulfoxide (DMSO)/H2O
- TNE buffer
- Detailed protocols (FluoReporter Blue Fluorometric dsDNA Quantitation Kit)
Each kit provides sufficient reagents for assaying 2000 samples using a fluorescence microplate reader.
Measurements of PCR products can be taken during the linear portion of the amplification reactions, allowing accurate quantitation of templates. Several methods exist for real-time quantitation of PCR products, including fluorescence resonance energy transfer techniques using fluorescently labeled primers or molecular beacon. Identification of PCR products during the reaction can also be monitored using Invitrogen SYBR Green I nucleic acid gel stain (S7563, S7567, S7585); this method has been shown to be more precise than TaqMan assays using labeled oligonucleotide probes.![]() In addition, individual DNA molecules have been detected with on-line capillary PCR coupled with laser-induced fluorescence detection by adding SYBR Green I stain to the reaction mixture.
In addition, individual DNA molecules have been detected with on-line capillary PCR coupled with laser-induced fluorescence detection by adding SYBR Green I stain to the reaction mixture.![]()
SYBR Green I stain binds preferentially to dsDNA, allowing accurate quantitation of double-stranded product in the presence of single-stranded oligonucleotide primers.![]() SYBR Green I stain is stable to the extremes of temperature required for PCR reactions and does not interfere with Taq polymerase. Improved specificity for quantitating desired products can be achieved by using SYBR Green I stain after the assay to measure the melting temperature of the products.
SYBR Green I stain is stable to the extremes of temperature required for PCR reactions and does not interfere with Taq polymerase. Improved specificity for quantitating desired products can be achieved by using SYBR Green I stain after the assay to measure the melting temperature of the products.![]() Double-stranded DNA with no base mismatches will show a higher melting temperature than the nonspecific templates that contain mismatches. Real-time quantitative PCR experiments can be carried out using instruments specialized for the application
Double-stranded DNA with no base mismatches will show a higher melting temperature than the nonspecific templates that contain mismatches. Real-time quantitative PCR experiments can be carried out using instruments specialized for the application ![]() or by quantitating amplification products manually at different time points.
or by quantitating amplification products manually at different time points.![]() Real-time quantitative PCR with SYBR Green I stain has been used to develop reliable and simple assays for detecting genetic mutations, including duplications and deletions in mosquito drug-resistance genes,
Real-time quantitative PCR with SYBR Green I stain has been used to develop reliable and simple assays for detecting genetic mutations, including duplications and deletions in mosquito drug-resistance genes,![]() chromosomal translocations in human disease genes,
chromosomal translocations in human disease genes,![]() and base substitutions.
and base substitutions.![]() It has also been used for the unequivocal identification of viral, bacterial or fungal pathogens.
It has also been used for the unequivocal identification of viral, bacterial or fungal pathogens.![]() In addition, this method has been used successfully for quantitative reverse-transcription PCR.
In addition, this method has been used successfully for quantitative reverse-transcription PCR.![]()
Invitrogen has introduced the next generation of real-time PCR detection reagents, including the Invitrogen SYBR GreenER qPCR reagent system, which produces the extremely reliable gene expression data from real-time quantitative PCR. SYBR GreenER reagent is a novel dsDNA-binding dye that exhibits a brighter signal and significantly reduced PCR inhibition compared to the original SYBR Green dye, but nearly identical spectral characteristics, More information about the SYBR GreenER qPCR reagent system and the latest developments in real-time PCR detection can be found at the Real-Time PCR (qPCR) Learning Center.
For Research Use Only. Not for use in diagnostic procedures.