Search
Selecting the Right Reactive Fluorescent Dye
First, it’s important to identify the reactivity—the chemical group to which the dye will attach—of your molecule, to determine which reactive moieties on the dye are at your disposal (Table 1). For instance, antibodies, proteins, and peptides have primary amines (Figure 1A) and sulfhydryl groups that are readily labeled. Low molecular weight compounds often lack these reactivities, and other groups, such as aldehydes and ketones, are therefore used in labeling reactions (Figure 1B). Some molecular classes, such as synthetic oligonucleotides and peptides, can be synthesized with groups designed for fluorescent labeling; sulfhydryl groups are often incorporated into oligos to enable facile labeling with maleimide reactive moieties. Free thiols, although not very common in proteins, can be modified by thiol-reactive reagents (Figure 1C) for site-specific labeling, which can be especially important for labeling small proteins without disrupting their activity.
| Reactivity | Molecule Class | Reactive Moiety | Product Example |
|---|---|---|---|
| Amines | Antibodies, proteins, and peptides | Activated esters (NHS, SDP, TFP) | Alexa Fluor® 488 succinimide (NHS) ester |
| Thiols | Antibodies, proteins, and peptides | Maleimides | Alexa Fluor® 594 maleimide |
| Aldehydes, ketones | Low MW molecules (drugs, steroid hormones, reduced sugars, metabolic intermediates) | Hydrazides | Alexa Fluor® 647 hydrazide |
 | Figure 1. Labeling reactions of (A) a primary amine with a succinimidyl (NHS) ester, (B) an aldehyde or ketone with a hydrazine derivative, and (C) a thiol with a maleimide. |
Preparing the Optimal Conjugate
Next, it’s critical to consider the optimal labeling scenario. Although conjugating dyes to biomolecules is usually rather easy, preparing the optimal conjugate may require extensive experimentation. The preferred bioconjugate should generally be highly fluorescent—that is, have a high degree of labeling (DOL)—yet retain key functional parameters such as solubility, selective binding to a receptor or nucleic acid, activation or inhibition of a particular enzyme, or the ability to incorporate into a biological membrane. However, conjugates with the highest DOL often precipitate out of solution or bind nonspecifically. To preserve function or binding specificity, the optimal fluorescent conjugate may therefore need to have a less-than-maximal fluorescence yield. This consideration applies particularly to macromolecules, such as antibodies and proteins, with a large number of primary amines. For a synthetic oligo with just one amine or sulfhydryl “handle” per molecule, you can at the outset plan the reaction to achieve maximum yield (Figure 2).
The Alexa Fluor® series of dyes, which span the near-UV, visible, and near-IR spectrum, are exceptionally bright and also exhibit significantly less self-quenching of proximal fluorophores than most other dye classes. At a given DOL, these properties synergistically contribute to maximal fluorescence of your biomolecule conjugate.
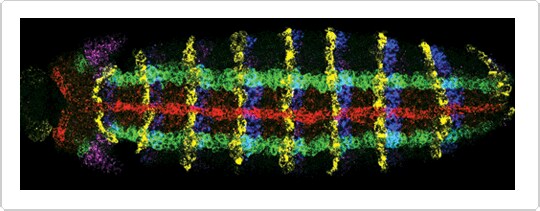
Figure 2. Simultaneous detection of expression of five genes in a whole-mount Drosophila embryo by Fluorescence in situ hybridization (FISH )with five RNA probes. Red: sog labeled using aminoallyl UTP and Alexa Fluor® 647 succinimidyl ester. Green: ind labeled with DNP, followed by rabbit anti–dinitrophenyl-KLH IgG antibody prelabeled with the Zenon® Alexa Fluor® 555 Rabbit IgG Labeling Kit. Blue: en labeled with biotin and detected with HRP–streptavidin and Alexa Fluor® 405 tyramide. Yellow: wg labeled with digoxigenin and detected with sheep anti–digoxigenin IgG antibody and Alexa Fluor® 594 donkey anti–sheep IgG antibody. Magenta: msh labeled with fluorescein and detected with mouse anti–fluorescein/Oregon Green® IgG2a antibody and Alexa Fluor® 488 goat anti–mouse IgG antibody. Image contributed by Dave Kosman and Ethan Bier, University of California, San Diego.
Choosing a Fluorescent Labeling Product
Some of the more commonly used fluorophores are listed in Table 2. Typical labeling reactions generally require the reactive fluorophore to first be dissolved in anhydrous dimethyl formamide (DMF) or dimethyl sulfoxide (DMSO). However, an advantage of Alexa Fluor® dyes is that their high aqueous solubility enables dissolution of the dye in buffer. For initial experiments using this approach, we recommend our proprietary Alexa Fluor® 488 5-SDP ester; SDP esters are considerably more stable toward hydrolysis than NHS esters, giving you better control over the labeling reaction. The dye solution is then added at a given molar ratio to the biomolecule dissolved in a suitable buffer. Typically, carbonate or borate buffers (pH ~8.5) are used. Note that some common buffers (e.g., Tris and glycine) contain amine groups and thus should not be used for labeling proteins.
To learn more about our selection of convenient protein labeling kits for diverse applications, read the article about APEX® Labeling Kits. To find the optimal fluorophore for your experimental application, use our web-based Labeling Chemistry Selection Tool. For more information on fluorescent labeling and detection, see The Molecular Probes® Handbook, or visit our web resource on Molecular Probes® labeling chemistry.
Table 2. Examples of Reactive Fluorophores.
| Fluorophore * | Excitation Maximum (nm) | Emission Maximum (nm) | Succinimidyl Ester (Amine-Reactive) | Maleimide (Thiol-Reactive) | Hydrazide (Aldehyde- and Ketone-Reactive) |
|---|---|---|---|---|---|
| Alexa Fluor® 350 | 346 | 422 | A10168 | A30505 | A10439 |
| Alexa Fluor® 488 | 495 | 519 | A20000 | A10254 | A10436 |
| Alexa Fluor® 555 | 555 | 565 | A20009 | A20346 | A20501MP |
| Alexa Fluor® 568 | 578 | 603 | A20003 | A20341 | A10437 |
| Alexa Fluor® 594 | 590 | 617 | A20004 | A10256 | A10438 |
| Alexa Fluor® 647 | 650 | 668 | A20006 | A20347 | A20502 |
| Alexa Fluor® 680 | 682 | 702 | A20008 | A20344 | — |
| BODIPY® FL | 505 | 513 | D2184 | B10250 | D2371 |
| Fluorescein | 494 | 518 | C1311 | F150 | C356 |
| Pacific Blue™ | 410 | 455 | P10163 | P30506 | — |
| Pacific Orange™ | 404 | 553 | P30253 | P30507 | — |
| Tetramethylrhodamine | 555 | 580 | C1171 | T6027 | — |
| Texas Red® | 595 | 615 | T6134 | T6008 | T6256 |
* To help you choose the optimal reactive fluorophore, explore our Labeling Chemistry Selection Tool.
For Research Use Only. Not for use in diagnostic procedures.
See a complete listing of the products discussed in this article.
View products
- Learn More about Fluorescent Labeling and Detection
- Learn More about Molecular Probes® Labeling Chemistry
Get a copy of this article as it appears in the print version of BioProbes 63.
